Research Overview
Research in the Denes lab focuses on bacteriophage-based applications for food safety and whole genome sequencing for improved surveillance and outbreak detection of food-borne pathogens. We employ genomic approaches, such as whole genome sequencing and RNA sequencing, as well as traditional microbiology and genetics methodologies.
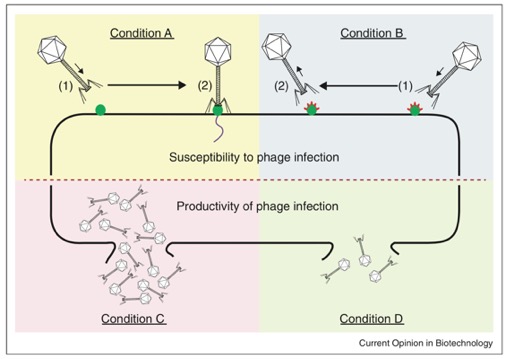
Bacteriophages (“phages”) are viruses that infect bacteria. We study the interactions between bacteriophages and bacterial food-borne pathogens to generate knowledge that can be used to develop improved phage-based applications. We are particularly interested in how bacterial pathogens develop resistance against phage infection. By understanding this process we can identify strategies to limit the emergence of phage resistance in environments where phage-based biocontrols or detection assays are used.
Whole genome sequencing (WGS) of recovered food-borne pathogen isolates is now routinely conducted to identify potential outbreaks. The Denes lab works with the Tennessee Department of Health (as part of the CDC-funded Tennessee Integrated Food Safety Center of Excellence) to improve our ability to incorporate WGS data in epidemiological investigation. We are also interested in how this data can be used to better understand the specific pathogen population structures responsible for food-borne illness in Tennessee.
Lab Members
- Lauren Hudson, postdoctoral researcher
- Education: University of Florida (BS), University of Georgia (PhD)
- Research Focus: Lauren uses computational tools to analyze whole genome sequencing data from food-borne pathogens isolated in Tennessee. She is determining their population structure, identifying unique mobile genetic elements, and identifying how to best coordinate the data with epidemiological investigations to identify potential outbreaks.
- Tracey Peters, PhD student
- Education: East Tennessee State University (BS)
- Research Focus: Tracey is performing coevolution studies between Listeria monocytogenes and its bacteriophages. She uses whole genome sequencing data and computational tools to identify mutations occurring in phage and host populations.
- Danielle Trudelle, MS student
- Education: State University of New York in Plattsburgh (BS)
- Research Focus: Danielle is examining cross-resistance to phage infection in Listeria monocytogenes. She is also working on new methods of determining cell envelope characteristics of different Listeria strains.
- Daniel Bryan, research specialist III
- Education: The Evergreen State College (BS)
